
To test the usability of our app, we applied TimeNexus together with PathLinker or ANAT on temporal expression data of the yeast cell cycle and were able to identify active subnetworks relevant for different cell cycle phases. We combined TimeNexus with the Cytoscape apps PathLinker and AnatApp/ANAT to extract active subnetworks from tMLNs. 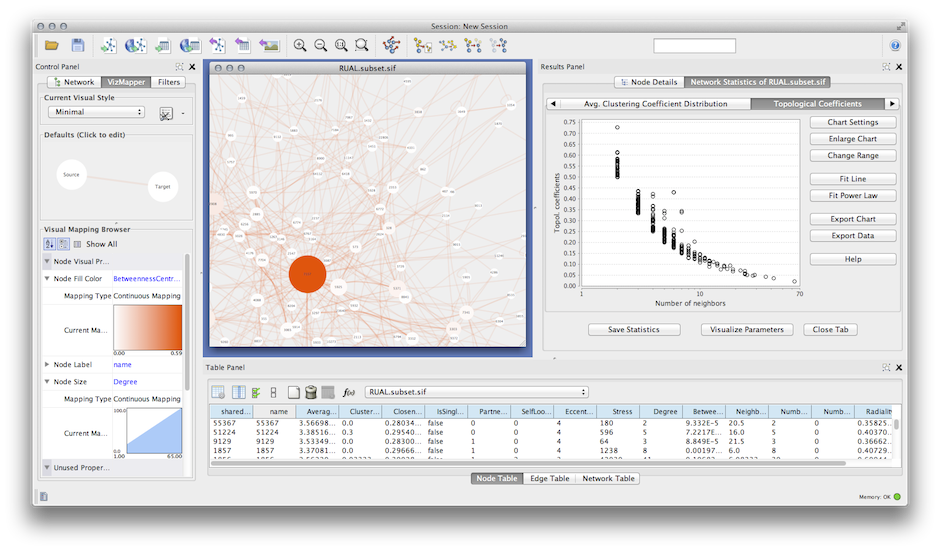
To allow further analysis of the tMLN, TimeNexus creates and passes on regular Cytoscape networks in form of static versions of the tMLN in three different ways: (i) over the entire set of layers, (ii) over two consecutive layers at a time, (iii) or on one single layer at a time. TimeNexus allows to easily create, manage and visualize temporal multilayer networks starting from a combination of node and edge tables carrying the information on the temporal network structure. We implemented this method as a Cytoscape app we named TimeNexus. Here, we present a method to project time-series data on sequential layers of a multilayer network, thus creating a temporal multilayer network (tMLN). Determining regulations or exploring the network structure over time requires time-dependent networks which incorporate time as one component in their structure. Yet, most -omics data integration tools are restricted to static networks and therefore cannot easily be used for analyzing time-series data. Integrating -omics data with biological networks such as protein–protein interaction networks is a popular and useful approach to interpret expression changes of genes in changing conditions, and to identify relevant cellular pathways, active subnetworks or network communities.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
